Description:

Innovators at Michigan State University (MSU) have developed a novel method for dynamic tracking of proteins in living cells. MSU's novel method tags proteins with a wide variety of colors, enabling several different proteins to be detected and tracked simultaneously. Using this method, multiple color variants are genetically tagged onto a protein expressed in cells, allowing proteins to be visualized over a very broad light spectrum -- including bright fluorescents -- as soon as they are synthesized.
Unlike green fluorescent protein (GFP), which is expensive and requires significant activation time, MSU's low-cost technology allows proteins to be detected instantaneously. The ability to track specific proteins as they are expressed can provide critical tools for pharmaceutical research and development, genetic engineering as well as a myriad of cell and molecular biology experiments.
Download PDF
Benefits
- Unlimited color variants: Virtually all colors and fluorescent properties can be obtained and co-expressed with a gene of interest. Absorbance and light transmission of the chromophore ligand can be modulated across the visible range to near-infrared range (425 - 644 nm).
- Extremely fast: Colors become visible within minutes.
- Excellent biochemical properties: The fusion domain has an excellent combination of properties, including small size and compact domain (~16 kD), high affinity for ligands (nm dissociation constant), highly stable chromophore (>1 week at room temperature), and very high recombinant expression. Protein variants are available that are stable at pH levels lower than 1.8.
- Allows for temporal control of visualization: Since the fusion protein is only fluorescent upon the addition of the fluorophore, and the fusion binds the fluorophore within seconds, proteins present within a narrow time window can be specifically visualized.
- Multi-purpose: This technology can simultaneously tag different proteins and measure pH in one cell in different cellular locations, indicating protein location and environment. Proteins have been developed covering a broad range of pH (1.5 - 8.8 pH units).
- Functions in anaerobic environments: Unlike GFP, this technology does not require molecular oxygen to become fluorescent.
Applications
- Pharmaceutical: Developing new drugs (e.g., high-throughput and high-content screening assays)
- Medical: Pursuing new treatments and diagnostics for a variety of diseases
- Agriculture: Developing transgenic organisms and new crops
- Brain research: Analyzing brain circuitry
- Infectious disease: Researching viral entry and infection of viruses
- Cell and molecular biology:
- Viral infection
- Intra/Extracellular receptor-mediated binding
- Protein expression, trafficking, post-translational modifications, and targeted degradation
- Cellular movement, attachment, growth, programmed death, and homeostasis
Technology Details
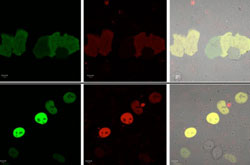 Figure 1. Demonstration of the CRBPII fluorescent protein in cells. (Top) The KLVF-eGFP fusion which is not localized. (Bottom) The KLVF mutant of CRBP II fused to eGFP (enhanced GFP) and Rb (retinoblastoma protein, which localizes to the nucleus (KLVF-eGFP-Rb).
Figure 1. Demonstration of the CRBPII fluorescent protein in cells. (Top) The KLVF-eGFP fusion which is not localized. (Bottom) The KLVF mutant of CRBP II fused to eGFP (enhanced GFP) and Rb (retinoblastoma protein, which localizes to the nucleus (KLVF-eGFP-Rb). Figure 2. Example of different color expression
Figure 2. Example of different color expression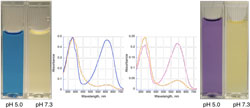 Figure 3. Example of use as a pH sensor
Figure 3. Example of use as a pH sensorThis technology for real-time imaging in live cells provides a low-cost alternative to existing fluorescent proteins, such as green fluorescent protein. The technique also can be used to determine real-time intracellular pH in discrete intracellular locations, organelles, and cellular structures.
The colorimetric or fluorescent tags can be used in a wide variety of recombinant and assay platforms to enable many research efforts, including cell-based protein localization, multi-color multiplex assays, in vitro detection, purification based on colorimetric characteristics, universal conjugation dyes, transgene selection markers, and natural food and plant colorings.
How It Works
Certain proteins can tolerate a wide variety of mutations in the ligand binding pocket. This is especially the case for the cellular retinoic acid-binding protein II (CRABP-II) and the human cellular retinol-binding protein II (hCRBPII) as well as other members of the intracellular lipid-binding protein (iLBP) family. Re-engineering of the proteins allows for binding of a variety of ligands with high affinity, and the resulting novel protein/ligand complexes exhibit a wide variety of colors encompassing the visible spectrum. When bound to an appropriate ligand, fluorescent proteins are also obtained.
Fluorescent tags can be detected using fluorescence microscopy or cytometry, and colorimetric dyes can be detected visually or using absorbance spectroscopy. By modulating the chromophore ligand, researchers can visualize virtually any color across the visible and near-infrared spectrum. The biochemical and colorimetric properties of these protein domains make them ideal as fusion tags in numerous research and development activities.
Why It Is Better
Unlike GFP, which requires several hours to manifest fluorescence, this protein fusion is fluorescent within minutes after the fluorophore has been added, meaning that proteins can be visualized as soon as they are synthesized. Furthermore, their visualization can be temporally controlled (i.e. proteins present at a single time point can be specifically visualized). MSU's technology is also remarkably stable, retaining its fluorescence at room temperature for several days in normal light. Because this method produces fluorescent proteins that are excited and emit over a broad range, several proteins can be detected simultaneously. This same system can be used for colorimetric and fluorescent detection of pH over a wide range. In addition, the method can be used in scenarios where GFP may be unsuitable (e.g., low intracellular oxygen environments, pulse-chase type experiments, low pH environments).
Publications/Patents
Michigan State University has applied for patent protection for this technology.
Additional Information
Commercial Opportunity
MSU Technologies (MSUT) facilitates the commercial development and public use of innovations developed by MSU faculty and staff. MSUT invites companies to consider opportunities to license this technology or engage in joint development activities.
About Us
The MSU Innovation Center seamlessly integrates three critical activities necessary to drive cutting edge innovations to market. The unit includes:
- MSU Technologies, reinvented with dynamic new leadership to make MSU's process for licensing and commercializing MSU intellectual property more productive and efficient;
- Business-CONNECT, formed to facilitate MSU's research relationships with the business community, including negotiating sponsored research agreements "at the speed of business"; and
- Spartan Innovations, a new unit that provides faculty and students with access to business talent, as well as operational and financial support to turn MSU innovations into successful Michigan businesses -- and spur entrepreneurship in the Michigan economy.
Co-located with East Lansing's Technology Innovation Center (TIC), and the Hatch, the MSU student company incubator, the MSU Innovation Center is the "center of gravity" for MSU's interactions with the private sector and support for entrepreneurism.